Harnessing the body’s immune system to treat cancer has made remarkable cures possible—but not for every patient. Now, researchers from UConn Health and the Cleveland Clinic have designed a way to reliably predict who will get good results from immunotherapy, they report in the Journal of Experimental Medicine.
Melanoma, a type of skin cancer, is often deadly once it spreads beyond its original spot. But immunotherapy drugs such as Keytruda have made dramatic cures possible, even for patients whose cancer has spread. About a third see their cancers stop growing or shrink, and 10% have complete remission. The flip side is that two-thirds of patients get no benefit at all.
“These immunotherapies are not really as effective as they seem in the advertisements” by drug companies, says UConn School of Medicine immunologist Beiyan Zhou. And the treatments take time. It can take at least two to four months, and as long as a year, to tell if a patient’s cancer responds to a specific immunotherapy. And if it doesn’t, that’s four months to a year of additional time to metastasize, when the patient could have been pursuing a different treatment entirely.
Zhou has been working with doctors at the Cleveland Clinic to figure out who will benefit from a specific immunotherapy before they spend time and money on it.
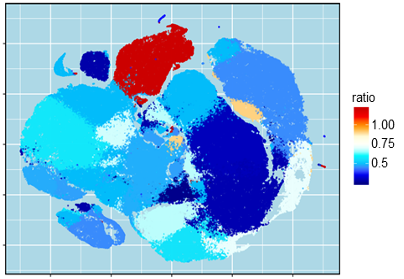
“Our study highlights a population of cells that not only is associated with immunotherapy resistance, but its intrinsic genetic signals can be leveraged to identify who will or will not respond to their emerging cancer therapeutics. Collaborative efforts of clinicians, immunologists and bioinformaticians yield these exciting discoveries,” says Brian Gastman, a collaborator on the project and the Director of Melanoma & High Risk Skin Cancer Program at Cleveland Clinic.
Gastman had been taking skin cancer samples and blood samples from patients and performing single cell profiling with them. These data were then analyzed at Zhou’s lab at UConn Health, first using conventional methods to take a lone cell and profile which genes in the cell are active.
Most cancer drug prediction tools rely on genetic analysis of specific mutations. If those mutations are present in the cancer, the tool will predict a certain drug will be effective. But these predictions are often inaccurate. Sometimes the gene with the mutation in question isn’t active or doesn’t matter to that particular cancer.
Unlike genetic analysis, single cell analysis tell us which genes a cell is actually using. Drugs that target active genes should be more effective than those targeting inactive ones.
Zhou and colleagues at UConn created a tool that used both computational and biomedical models on single cell data to predict whether or not a specific patient would respond to immunotherapy. After identifying a signature gene panel using a new strategy developed in Zhou’s lab, a predictive model was trained using a published single cell dataset (from Broad Institute of MIT and Harvard), and then tested in additional data from the Cleveland Clinic’s patient data. The tool, named NiCir (non-invasive circulating T cells model), focuses on the activity of 20 specific genes known to be important in melanoma skin cancer and lung cancer. NiCir achieved 88% accuracy in three different datasets predicting which patient’s cancers had shrunken in response to a specific immuno-checkpoint inhibitor immunotherapy.
The team is now working on creating an new computational program which will allow the creation of more accurate tools for disease prediction, and to facilitate guided-drug discovery.